Home
Research
Our group investigates the biology of the bacterial cell envelope. In Gram-negative bacteria this structure comprises two membranes (the cytoplasmic and outer membranes) and the periplasmic compartment between them which contains the cell wall. We are particularly interested in understanding how proteins and DNA are transported across and around the cell envelope.
We address these problems using a very wide range of techniques (some collaboratively) including protein biochemistry, molecular genetics, bacterial cell biology, single molecule fluorescence imaging, structural biology, and molecular simulations.
Our work is funded by Wellcome and the European Research Council.

The Unusual Cell Envelope of Bacteroidota Bacteria
Bacteroidota are the most abundant Gram-negative commensal bacteria in the human gut, where they play essential roles in maintaining gut health. They are also major constituents of other human microbiomes. The Bacteroidota include major human opportunistic anaerobic pathogens responsible for sepsis and severe dental disease (periodontitis). Beyond the human body, Bacteroidota are environmentally widespread and are key contributors to the microbial biosphere and plant health.
Recent research has revealed that the biology of the Bacteroidota cell envelope is very different from that found in the well-characterised bacterial model Escherichia coli. Our work focuses on investigating the most important of these differences.
Protein transport by the Bacteroidota-specific Type 9 Secretion System
Pathogenic bacteria use protein secretion systems to export effector proteins that manipulate their host and promote infection. The Bacteroidota make only limited use of the protein export pathways found in other phyla but instead use a novel, phylum-specific Type 9 Secretion System (T9SS) to deliver proteins into their extracellular environment.
The T9SS is the most complex protein transport system known. The T9SS contains at least 20 protein components, the functions of which include forming an enormous regulated pore in the outer membrane (the translocon), modifying substrate proteins after transport, and energization of transport using the proton motive force across the cytoplasmic membrane. We study all parts of the pathway with the aim of elucidating the molecular mechanism of Type 9 transport.
Our research in this area is sponsored by the ERC (SecNine ERC-2018-ADG H2020 833713) and Wellcome (Discovery Award 317933/Z/24/Z).
Structure of a substrate-bound Type 9 translocon complex. The channel protein (blue) forms a ~70 Å diameter transmembrane pore. A carrier protein (brown) inserts through the extracellular exit of the pore to bind the targeting signal (CTD) of the substrate protein (red) within the pore lumen. Subsequent dissociation of the carrier protein from the translocon pulls the bound substrate protein through the channel pore. For more information see Lauber et al. (2024) Nat Microbiol 9: 1089–1102.

The T9SS is energised by a trans-envelope energy chain. The proton motive force across the inner membrane (IM) drives proton flow through Motor complexes, inducing rotation of their periplasmic rotor arms. These rotating Motor arms in turn drive rotation of the circular PorKN complex located at the outer membrane (OM). Rotation of PorKN supplies the mechanical energy to power protein transport through the T9SS translocon. Experimental structures are shown in spacefilling representation. For more information see Liu et al. (2025) Nature Comms 16: 10217.
Recent publications
Liu, X., Avramova, M., Deme, J.C., Jones, R.L., Lundgren, C.A.K., Lea, S.M., and Berks, B.C. (2025) A shared mechanism for Bacteroidota protein transport and gliding motility. Nature Comms 16: 10217. https://www.nature.com/articles/s41467-025-65003-8
Lauber, F., Deme, J.C., Liu, X., Kjær, A., Miller, H.L., Alcock, F., Lea, S.M., and Berks, B.C. (2024) Structural insights into the mechanism of protein transport by the Type 9 Secretion System translocon. Nat Microbiol 9: 1089–1102. https://www.nature.com/articles/s41564-024-01644-7
Hennell James, R., Deme, J.C., Kjær, A., Alcock, F., Silale, A., Lauber, F., Johnson, S., Berks, B.C., and Lea, S.M. (2021) Structure and mechanism of the proton-driven motor that powers type 9 secretion and gliding motility. Nat Microbiol 6: 221–233. https://www.nature.com/articles/s41564-020-00823-6
Lauber, F., Deme, J.C., Lea, S.M., and Berks, B.C. (2018) Type 9 secretion system structures reveal a new protein transport mechanism. Nature 564: 77–82. https://www.nature.com/articles/s41586-018-0693-y
Bacteroidota gliding motility
Motile Bacteroidota move by gliding across solid surfaces using a mechanism unique to this phylum. This allows Bacteroidota bacteria to travel at speeds hundreds of times faster than eukaryotic cells. Gliding motility is mediated by the helical movement of adhesin molecules along the cell surface. We aim to understand the molecular basis of this process and how motility is regulated in response to external stimuli.
Our recent work has uncovered a close mechanistic and evolutionary relationship between the Bacteroidota gliding apparatus and the T9SS. Based on these findings, we propose that the T9SS is a miniaturized version of the machinery that powers cell movement in gliding Bacteroidota.
Our research in this area is sponsored by the ERC (SecNine ERC-2018-ADG H2020 833713) and Wellcome (Discovery Award 317933/Z/24/Z).
A - Bacteroidota bacteria gliding on glass imaged in real time.
B - Gliding cells move using a corkscrew motion. Cell motion is revealed by imaging a fluorescently labelled protein that has a fixed position on the cell surface. The movie plays in real time. From Lauber et al. (2018) Nature 564: 77–82.
C - Gliding adhesins move in helical trajectories on the cell surface. A single fluorescently labelled adhesin molecule is imaged in real time moving along the surface of an immobilised cell, with the super-resolution localizations of its trajectory plotted.
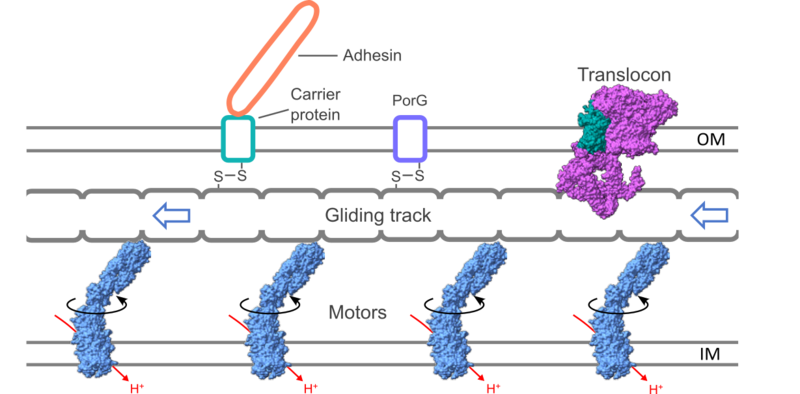
The organization of the gliding motility apparatus is related to that of the T9SS. The mobile gliding track is structurally and functionally analogous to the PorKN ring of the T9SS and is likewise propelled by protonmotive force-driven Motors. The cell-surface gliding adhesins (orange) are connected to the gliding track by a carrier protein (sea green) in the outer membrane, which is disulfide-linked to the track. Experimental structures are shown in spacefilling representation. From Liu et al. (2025) Nature Comms 16: 10217.
Recent publications
Liu, X., Avramova, M., Deme, J.C., Jones, R.L., Lundgren, C.A.K., Lea, S.M., and Berks, B.C. (2025) A shared mechanism for Bacteroidota protein transport and gliding motility. Nature Comms 16: 10217. https://www.nature.com/articles/s41467-025-65003-8
Hennell James, R., Deme, J.C., Kjær, A., Alcock, F., Silale, A., Lauber, F., Johnson, S., Berks, B.C., and Lea, S.M. (2021) Structure and mechanism of the proton-driven motor that powers type 9 secretion and gliding motility. Nat Microbiol 6: 221–233. https://www.nature.com/articles/s41564-020-00823-6
Outer membrane protein biogenesis in the Bacteroidota
Gram-negative bacteria are distinguished by the presence of an outer membrane (OM) at the cell periphery. This membrane is the site at which the bacterium interacts with its environment (or with its host if a pathogen), and provides the first line of defence against antibiotics, mechanical stress, and immunological attacks. These functions depend on proteins that either span the outer membrane (Outer Membrane Proteins, or OMPs) or are anchored to it by a lipid tail (lipoproteins). Sophisticated machinery is required to target, insert, and fold these proteins. This machinery is well-characterised in the model bacterium E. coli. However, Bacteroidota possess OM proteins that differ significantly from those typically found in the E. coli OM proteome, raising the question of how these proteins are assembled.
We recently discovered that the BAM complex that inserts OMPs into the OM has a very different composition in Bacteroidota relative to the E. coli paradigm, including the presence of novel extracellular subunits. We now seek to identify the functions of these additional subunits and to elucidate their molecular mechanism of action.
Our research in this area is sponsored by the ERC (SecNine ERC-2018-ADG H2020 833713 and BactOMPbuild ERC-2024-ADG HORIZON 101201241).

Bacteroidota OMPs are more complex than those found in E. coli. Bacteroidota OMPs (blue) characteristically have larger extracellular domains and there are many cell surface lipoproteins (salmon).

Bacteroidota BAM is more complex than E. coli BAM. BAM is the machine responsible for OMP insertion into the OM. Substrate OMPs fold at the lateral seam. Subunits found only in Bacteroidota BAM are shown in salmon. For more details see Liu, Orenday Tapia et al. (2025) Nature 647: 479–487.
Recent publications
Liu, X., Orenday Tapia, L., Deme, J.C., Lea, S.M., and Berks, B.C. (2025) A new paradigm for outer membrane protein biogenesis in the Bacteroidota. Nature 647: 479–487. https://www.nature.com/articles/s41586-025-09532-8
Mechanism of horizontal gene transfer between bacteria by conjugation
In conjugation a plasmid or other genetic element in a donor bacterium is transferred to a recipient bacterium through a mating bridge between the two cells. It is the main route for the spread of antibiotic resistance and other adaptive traits important for pathogens. Conjugation requires the transport of DNA molecules across the cell envelopes of both donor and recipient bacteria. We study the molecular mechanisms by which this occurs.
Structure of the conjugation exclusion protein TraT. Conjugal plasmids encode exclusion systems that prevent redundant plasmid transfer into cells already carrying the same plasmid. TraT is an outer membrane exclusion protein that blocks incoming conjugative pili at the recipient cell surface, thereby blocking DNA transfer. For more details see Chen et al. (2025) Communications Biology 8: 1702.
Recent publications
Chen, N., Bukys, A., Lundgren, C.A.K., Deme, J.C., El Sayyed, H., Kapanidis, A.N., Lea, S.M., and Berks, B.C. (2025) Structure of the conjugation surface exclusion protein TraT. Communications Biology 8: 1702 https://www.nature.com/articles/s42003-025-09102-8
Protein transport by the Tat System
The twin-arginine translocation (Tat) system is a mechanistically unique protein transport pathway moving folded proteins across membranes. It is found in all domains of life and is essential for plant photosynthesis and bacterial virulence.
A particular mechanistic challenge for the Tat system is that the diverse shapes and sizes of folded proteins make it difficult to maintain the membrane seal around the transporting protein. Our laboratory seeks to understand the molecular basis of this unusual transport system.

Schematic model of the Tat translocation cycle The resting state Tat receptor in E. coli comprises several copies of TatA, TatB and TatC (here only one copy of each is shown for clarity). (1) The signal peptide of a folded Tat substrate engages the receptor primarily through interaction with TatC. (2) This triggers recruitment of additional TatA protomers. The proton motive force (PMF) is required at this stage. (3) The folded substrate moves across the membrane, facilitated by the TatA oligomer. (4) Upon completion of translocation, the signal peptide is cleaved from the substrate protein, and the TatA oligomer disperses, resetting the receptor complex to its resting state. From Palmer and Berks (2024) Curr Biol 34: R267–R268.

Structure of a substrate-bound Tat core complex. For more information see Deme, Bryant et al. (2026) bioRxiv doi: https://doi.org/10.1101/2025.09.18.677151
Recent publications
Deme, J.C., Bryant, O.J., Batista, M.R.B., Stansfeld, P.J., Berks, B.C., and Lea, S.M. (2026) Structure and substrate recognition by the Twin-arginine translocation (Tat) pathway core complex. bioRxiv doi: https://doi.org/10.1101/2025.09.18.677151
Palmer, T., and Berks, B.C. (2024) The twin-arginine translocation (Tat) system. Curr Biol 34: R267–R268. https://www.cell.com/current-biology/fulltext/S0960-9822(24)00214-8
Some of our collaborators
- Susan Lea (structural biology) https://www.stjude.org/people/l/susan-lea.html
- Achillefs Kapanidis (single molecule fluorescence/conjugation) https://kapanidis.web.ox.ac.uk/home
- Richard Berry (bacterial motility) https://www.physics.ox.ac.uk/our-people/berryr
- Phillip Stansfeld (molecular simulations) https://warwick.ac.uk/fac/sci/lifesci/people/pstansfeld/


